Briggs and researchers at the Max Planck Institute for Biochemistry have discovered the mechanism behind a crucial step in the HIV life cycle. In collaboration with teams from Heidelberg University and Yale University, they revealed that the mysterious “spacer peptide 2” (one of the HIV viral components) plays a key role in transforming immature HIV-1 particles into infectious particles. The research results have been published in the journal Nature.
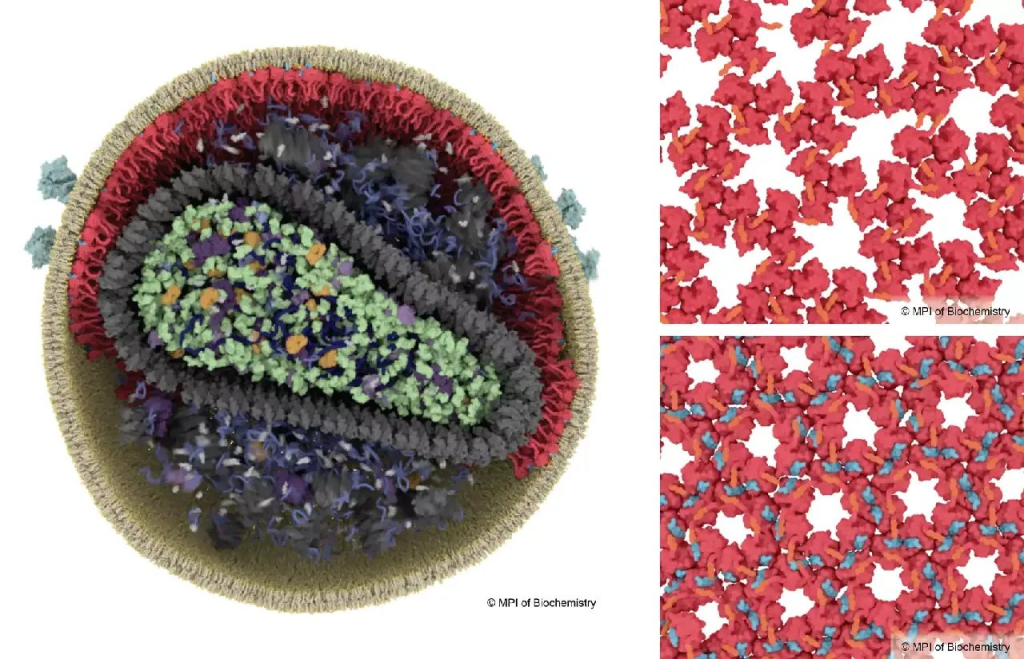
Key processes in HIV maturation
HIV needs to undergo a maturation process to become infectious, which involves rearrangements in the structure of the matrix protein (red). In immature virus particles, the matrix protein forms a loose lattice structure (top right), while in mature HIV, the binding of spacer peptide 2 triggers the formation of a new, more stable lattice structure (blue, bottom right). © Illustration: Margot Riggi, MPI of Biochemistry.
HIV-1 particle maturation
HIV-1 particles are initially released from infected cells in an immature, non-infectious form. The main building block of the virus particle is about 2,000 copies of a long rod-shaped protein called Gag. To be able to infect host cells, HIV must go through a maturation process that involves the cleavage of Gag by HIV-1 protease (a viral enzyme), breaking it down into six smaller proteins, including the capsid protein and the matrix protein. This cleavage process results in a huge rearrangement of the virus’s internal structure.
Although scientists have been exploring structural changes in the viral capsid that envelops the genome for many years, comparatively little is known about the virosome, the protein shell beneath the lipid membrane that envelops the virus. A research team led by structural biologist Johann Briggs, director of the Max Planck Institute of Biochemistry, has now solved the mystery of how the matrix proteins rearrange themselves during maturation into infectious particles.
New discoveries revealed using cryo-electron microscopy
The scientists used the latest cryo-electron microscopy technology to image the virus particles and derived a very detailed 3D model of the viral proteins by computational image analysis. To their surprise, the rearrangement of the matrix was triggered by “spacer peptide 2”, which binds to the matrix proteins and causes the matrix to stack up in a different way. Spacer peptide 2 is one of the six components formed by cleaved Gag, but its function has remained unclear until now. The binding of spacer peptide 2 to the matrix proteins enables the virus to fuse more quickly with the target cell. The study was conducted in collaboration with research teams from Heidelberg University and Yale University.
Researchers’ explanation and future prospects
John Briggs said: “In our lab, we obtained the first structural data on the matrix in 2021, but at that time it was not clear what caused it to rearrange as the virus matured. In this new study, we produced a more detailed 3D view of the matrix, which is crucial to understanding the changes that are taking place.”
The two first authors, James Stacey and Dominic Herbik, explain their findings. James says: “The virosome has a pocket in its mature form. We knew something was binding to it, but we thought it was membrane lipids. Now we see that it is spacer peptide 2. We can’t help but wonder if this pocket could be a target for drug molecules in the future.” Dominic adds: “Until now, the function of spacer peptide 2 has been unknown. Thanks to high-resolution cryo-electron microscopy, we can finally see that this peptide binds directly to the matrix proteins after viral release and links the proteins together in the mature virus.”
John Briggs concluded: “Although HIV-1 is probably one of the most studied viruses, we still don’t understand enough about some important steps in its replication process.”





